Matrix: Schulthess/N_biocarta
Description: biochemical network; left nullspace is required
 |
| (bipartite graph drawing) |
 |
 |
 |
| Matrix properties | |
| number of rows | 1,922 |
| number of columns | 1,996 |
| nonzeros | 4,335 |
| structural full rank? | no |
| structural rank | 1,185 |
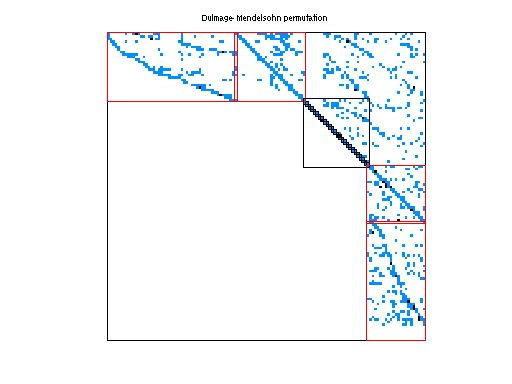
| # of blocks from dmperm | 347 |
| # strongly connected comp. | 800 |
| explicit zero entries | 0 |
| nonzero pattern symmetry | 0% |
| numeric value symmetry | 0% |
| type | integer |
| structure | rectangular |
| Cholesky candidate? | no |
| positive definite? | no |
| author | P. Schulthess |
| editor | T. Davis |
| date | 2012 |
| kind | biochemical network |
| 2D/3D problem? | no |
Notes:
Matrices from Pascal Schulthess, Institute for Pathology,
Chariteplatz 1, 10117 Berlin, Germany.
Three large biochemical networks (N_biocarta, N_pid, and N_reactome).
These are stoichiometric matrices extracted from three biochemical
databases (BioCarta, PID, and REACTOME) describing cell signaling
pathways and protein-protein interaction networks. The goal is to
find the left nullspace of the matrix; in MATLAB notation:
N = null (Problem.A') ;
The matrix (Problem.A')*N will thus be essentially zero.
This can be done much more efficiently with the spqr_rank toolbox by
Leslie Foster and Tim Davis, as:
N = spqr_null (Problem.A') ;
Results:
The matrix A is transposed, then N = null (A) or N = spqr_null (A)
is computed. The size statistic is the memory taken by N.
spqr_null can compute either an explicit matrix N, or an implicit
Householder-based representation. The latter takes less memory.
Matrix: N_biocarta size: 1996 by 1922 (transposed)
spqr_null stats:
flag: 0
rank: 1023
tol: 3.5456e-12
est_sval_upper_bounds: [0.1689 3.4534e-15]
est_sval_lower_bounds: [0.1203 0]
sval_numbers_for_bounds: [1023 1024]
est_norm_A_times_N: 2.4349e-15
spqr_null, implicit: 0.03 sec, norm(A*N) 9e-15 size: 0.08 MB
spqr_null, explicit: 0.10 sec, norm(A*N) 9e-15 size: 0.11 MB
MATLAB null: 3.31 sec, norm(A*N) 2e-13 size: 13.82 MB
all report dim(N) of 899.
Matrix: N_pid size: 3923 by 3625 (transposed)
spqr_null stats:
flag: 0
rank: 2048
tol: 1.3937e-11
est_sval_upper_bounds: [0.0922 5.1310e-15]
est_sval_lower_bounds: [0.0585 0]
sval_numbers_for_bounds: [2048 2049]
est_norm_A_times_N: 1.6751e-15
spqr_null, implicit: 0.05 sec, norm(A*N) 4e-14 size: 0.21 MB
spqr_null, explicit: 0.34 sec, norm(A*N) 4e-14 size: 1.32 MB
MATLAB null: 24.86 sec, norm(A*N) 9e-13 size: 45.73 MB
all report dim(N) of 1577
Matrix: N_reactome size: 16559 by 10204 (transposed)
spqr_null stats:
flag: 0
rank: 9025
tol: 1.1766e-10
est_sval_upper_bounds: [0.6722 1.3042e-14]
est_sval_lower_bounds: [0.0106 0]
sval_numbers_for_bounds: [9025 9026]
est_norm_A_times_N: 9.4695e-15
spqr_null, implicit: 0.95 sec, norm(A*N) 2e-13 size: 7.5 MB
spqr_null, explicit: 3.53 sec, norm(A*N) 2e-13 size: 25.2 MB
MATLAB null: 904.54 sec, norm(A*N) 2e-10 size: 96.2 MB
all report dim(N) of 1179.
| Ordering statistics: | result |
| nnz(V) for QR, upper bound nnz(L) for LU, with COLAMD | 7,931 |
| nnz(R) for QR, upper bound nnz(U) for LU, with COLAMD | 4,139 |
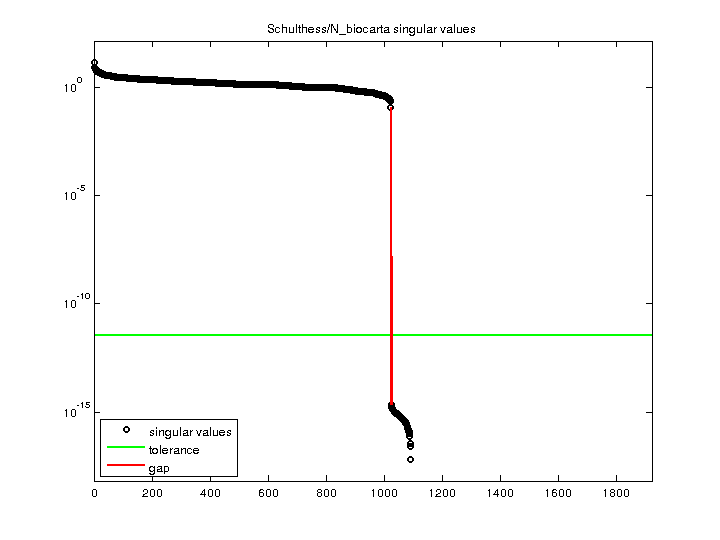
| SVD-based statistics: | |
| norm(A) | 14.8187 |
| min(svd(A)) | 0 |
| cond(A) | Inf |
| rank(A) | 1,023 |
| sprank(A)-rank(A) | 162 |
| null space dimension | 899 |
| full numerical rank? | no |
| singular value gap | 5.50622e+13 |
| singular values (MAT file): | click here |
| SVD method used: | s = svd (full (R)) ; where [~,R,E] = spqr (A') with droptol of zero |
| status: | ok |
For a description of the statistics displayed above, click here.
Maintained by Tim Davis, last updated 12-Mar-2014.
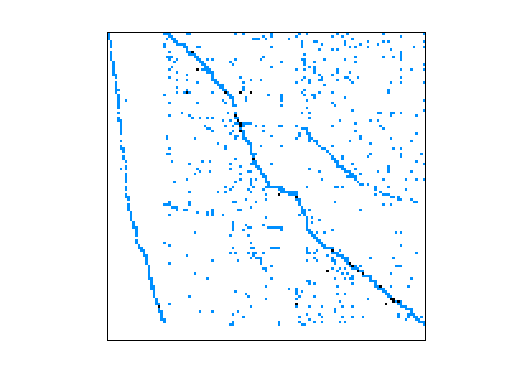
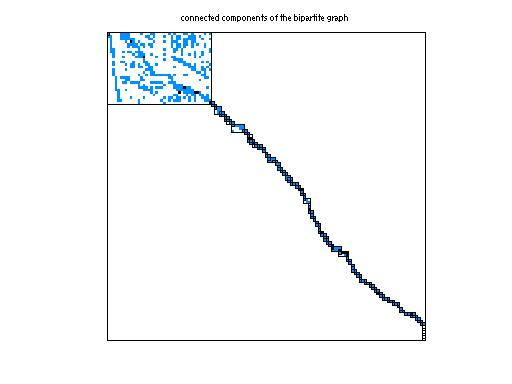
Matrix pictures by cspy, a MATLAB function in the CSparse package.
Matrix graphs by Yifan Hu, AT&T Labs Visualization Group.