Matrix: SNAP/ca-CondMat
Description: Collaboration network of Arxiv Condensed Matter
 |
| (undirected graph drawing) |
 |
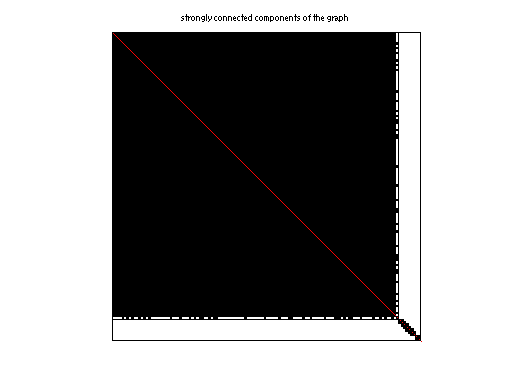 |
| Matrix properties | |
| number of rows | 23,133 |
| number of columns | 23,133 |
| nonzeros | 186,936 |
| # strongly connected comp. | 567 |
| explicit zero entries | 0 |
| nonzero pattern symmetry | symmetric |
| numeric value symmetry | symmetric |
| type | binary |
| structure | symmetric |
| Cholesky candidate? | no |
| positive definite? | no |
| author | J. Leskovec, J. Kleinberg and C. Faloutsos |
| editor | J. Leskovec |
| date | 2003 |
| kind | undirected graph |
| 2D/3D problem? | no |
| Additional fields | size and type |
| nodename | full 23133-by-1 |
Notes:
Networks from SNAP (Stanford Network Analysis Platform) Network Data Sets,
Jure Leskovec http://snap.stanford.edu/data/index.html
email jure at cs.stanford.edu
Condense Matter collaboration network
Dataset information
Arxiv COND-MAT (Condense Matter Physics) collaboration network is from the
e-print arXiv and covers scientific collaborations between authors papers
submitted to Condense Matter category. If an author i co-authored a paper with
author j, the graph contains a undirected edge from i to j. If the paper is
co-authored by k authors this generates a completely connected (sub)graph on k
nodes.
The data covers papers in the period from January 1993 to April 2003 (124
months). It begins within a few months of the inception of the arXiv, and thus
represents essentially the complete history of its COND-MAT section.
Dataset statistics
Nodes 23133
Edges 186936
Nodes in largest WCC 21363 (0.923)
Edges in largest WCC 182628 (0.977)
Nodes in largest SCC 21363 (0.923)
Edges in largest SCC 182628 (0.977)
Average clustering coefficient 0.6334
Number of triangles 173361
Fraction of closed triangles 0.2643
Diameter (longest shortest path) 15
90-percentile effective diameter 6.6
Source (citation)
J. Leskovec, J. Kleinberg and C. Faloutsos. Graph Evolution: Densification and
Shrinking Diameters. ACM Transactions on Knowledge Discovery from Data (ACM
TKDD), 1(1), 2007.
Files
File Description
ca-CondMat.txt.gz Collaboration network of Arxiv Condensed Matter category
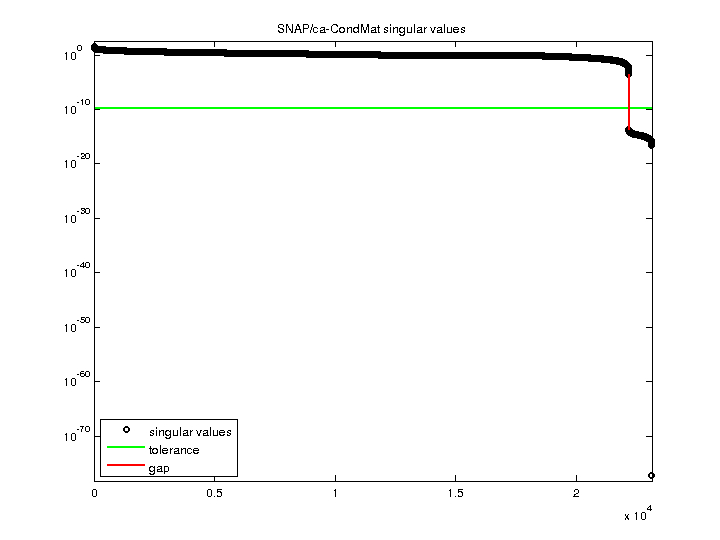
| SVD-based statistics: | |
| norm(A) | 37.9541 |
| min(svd(A)) | 5.52096e-78 |
| cond(A) | 6.87455e+78 |
| rank(A) | 22,177 |
| null space dimension | 956 |
| full numerical rank? | no |
| singular value gap | 1.26088e+10 |
| singular values (MAT file): | click here |
| SVD method used: | s = svd (full (A)) |
| status: | ok |
For a description of the statistics displayed above, click here.
Maintained by Tim Davis, last updated 12-Mar-2014.
Matrix pictures by cspy, a MATLAB function in the CSparse package.
Matrix graphs by Yifan Hu, AT&T Labs Visualization Group.