Matrix: SNAP/Oregon-2
Description: (9 graphs) AS peering info inferred from Oregon route-views, 3/31-5/26/01
 |
| (undirected graph drawing) |
 |
| Matrix properties | |
| number of rows | 11,806 |
| number of columns | 11,806 |
| nonzeros | 65,460 |
| # strongly connected comp. | 346 |
| explicit zero entries | 0 |
| nonzero pattern symmetry | symmetric |
| numeric value symmetry | symmetric |
| type | binary |
| structure | symmetric |
| Cholesky candidate? | no |
| positive definite? | no |
| author | J. Leskovec, J. Kleinberg and C. Faloutsos |
| editor | J. Leskovec |
| date | 2001 |
| kind | undirected graph sequence |
| 2D/3D problem? | no |
| Additional fields | size and type |
| G | cell 9-by-1 |
| Gname | full 9-by-14 |
| nodename | full 11806-by-1 |
Notes:
Networks from SNAP (Stanford Network Analysis Platform) Network Data Sets,
Jure Leskovec http://snap.stanford.edu/data/index.html
email jure at cs.stanford.edu
Autonomous systems - Oregon-2
Dataset information
9 Autonomous systems graphs, 1 per week between March 31 2001 and May 26 2001.
Graphs represent AS peering information inferred from Oregon route-views,
Looking glass data, and Routing registry, all combined.
Dataset statistics are calculated for the graph with the lowest (March 31 2001)
and highest (from May 26 2001) number of nodes:
Dataset statistics for graph with lowest number of nodes - 3 31 2001
Nodes 10900
Edges 31180
Nodes in largest WCC 10900 (1.000)
Edges in largest WCC 31180 (1.000)
Nodes in largest SCC 10900 (1.000)
Edges in largest SCC 31180 (1.000)
Average clustering coefficient 0.5009
Number of triangles 82856
Fraction of closed triangles 0.03855
Diameter (longest shortest path) 9
90-percentile effective diameter 4.3
Dataset statistics for graph with highest number of nodes - 5 26 2001
Nodes 11461
Edges 32730
Nodes in largest WCC 11461 (1.000)
Edges in largest WCC 32730 (1.000)
Nodes in largest SCC 11461 (1.000)
Edges in largest SCC 32730 (1.000)
Average clustering coefficient 0.4943
Number of triangles 89541
Fraction of closed triangles 0.03701
Diameter (longest shortest path) 9
90-percentile effective diameter 4.3
Source (citation)
J. Leskovec, J. Kleinberg and C. Faloutsos. Graphs over Time: Densification
Laws, Shrinking Diameters and Possible Explanations. ACM SIGKDD International
Conference on Knowledge Discovery and Data Mining (KDD), 2005.
Files
File Description
AS peering information inferred from Oregon route-views, Looking glass
data, and Routing registry, ...
oregon2_010331.txt.gz from March 31 2001
oregon2_010407.txt.gz from April 7 2001
oregon2_010414.txt.gz from April 14 2001
oregon2_010421.txt.gz from April 21 2001
oregon2_010428.txt.gz from April 28 2001
oregon2_010505.txt.gz from May 05 2001
oregon2_010512.txt.gz from May 12 2001
oregon2_010519.txt.gz from May 19 2001
oregon2_010526.txt.gz from May 26 2001
NOTE: for the UF Sparse Matrix Collection, the primary matrix in this problem
set (Problem.A) is the last matrix in the sequence, oregon2_010526, from May 26
2001.
The nodes are uniform across all graphs in the sequence in the UF collection.
That is, nodes do not come and go. A node that is "gone" simply has no edges.
This is to allow comparisons across each node in the graphs.
Problem.aux.nodenames gives the node numbers of the original problem. So
row/column i in the matrix is always node number Problem.aux.nodenames(i) in
all the graphs.
Problem.aux.G{k} is the kth graph in the sequence.
Problem.aux.Gname(k,:) is the name of the kth graph.
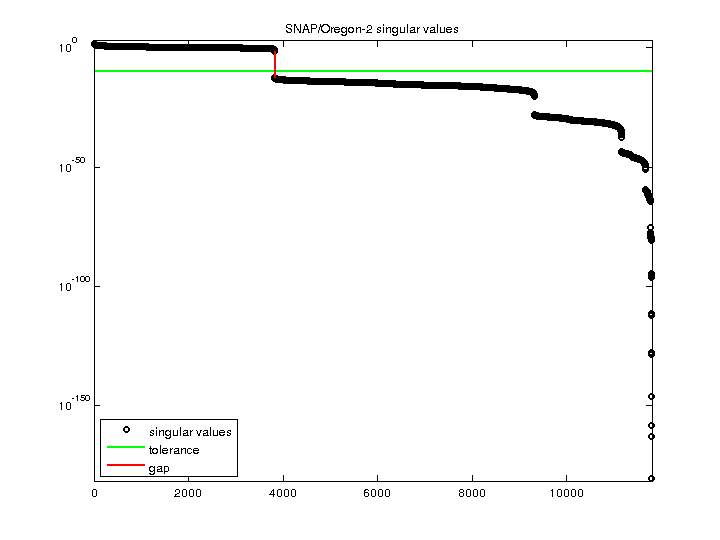
| SVD-based statistics: | |
| norm(A) | 75.2407 |
| min(svd(A)) | 0 |
| cond(A) | Inf |
| rank(A) | 3,825 |
| null space dimension | 7,981 |
| full numerical rank? | no |
| singular value gap | 9.24486e+10 |
| singular values (MAT file): | click here |
| SVD method used: | s = svd (full (A)) ; |
| status: | ok |
For a description of the statistics displayed above, click here.
Maintained by Tim Davis, last updated 12-Mar-2014.
Matrix pictures by cspy, a MATLAB function in the CSparse package.
Matrix graphs by Yifan Hu, AT&T Labs Visualization Group.