Matrix: Pajek/EAT_RS
Description: Pajek network: Edinburgh Associative Thesaurus (response-stimulus)
 |
 |
| (bipartite graph drawing) | (graph drawing of A+A') |
 |
 |
| Matrix properties | |
| number of rows | 23,219 |
| number of columns | 23,219 |
| nonzeros | 325,592 |
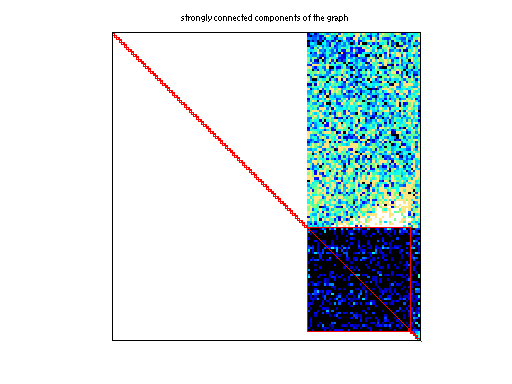
| # strongly connected comp. | 15,466 |
| explicit zero entries | 0 |
| nonzero pattern symmetry | 12% |
| numeric value symmetry | 3% |
| type | integer |
| structure | unsymmetric |
| Cholesky candidate? | no |
| positive definite? | no |
| author | G. Kiss, C. Armstrong R. Milroy, J. Piper |
| editor | V. Batagelj |
| date | 1971 |
| kind | directed weighted graph |
| 2D/3D problem? | no |
| Additional fields | size and type |
| nodename | full 23219-by-20 |
Notes:
------------------------------------------------------------------------------
Pajek network converted to sparse adjacency matrix for inclusion in UF sparse
matrix collection, Tim Davis. For Pajek datasets, See V. Batagelj & A. Mrvar,
http://vlado.fmf.uni-lj.si/pub/networks/data/.
------------------------------------------------------------------------------
EAT - The Edinburgh Associative Thesaurus /
response-stimulus
--------------------------------------------------------
The EAT is a database of word association norms.
- Original EAT: George Kiss, Christine Armstrong,
Robert Milroy and J.R.I. Piper (1968-1971).
- MRC Psycholinguistic Database Version modified by:
Max Coltheart, S. James, J. Ramshaw, B.M. Philip,
B. Reid, J. Benyon-Tinker and E. Doctor;
made available by: Philip Quinlan.
- The present version was re-structured and documented
by Michael Wilson at the Rutherford Appleton Laboratory.
http://monkey.cis.rl.ac.uk/Eat/htdocs/eat.zip
transformed in Pajek format: V. Batagelj, 31. July 2003
-----
------------------------------------------------------------------------------
Regarding conversion for UF sparse matrix collection: in the original data
there are 325,624 weighted edges. Of those only 32 edges are duplicates, and
all of them have identical edge weights as the edges they are duplicates of
These extraneous edges have been removed, since this this appears to be a
graph, not a multigraph.
------------------------------------------------------------------------------
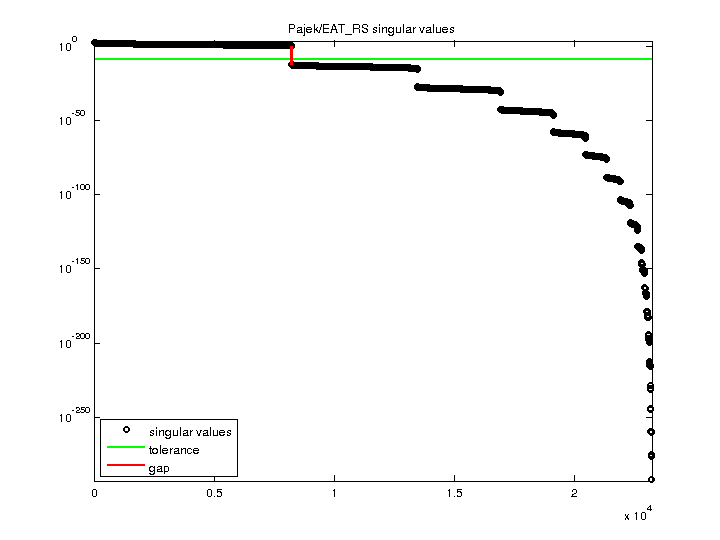
| SVD-based statistics: | |
| norm(A) | 312.442 |
| min(svd(A)) | 0 |
| cond(A) | Inf |
| rank(A) | 8,210 |
| null space dimension | 15,009 |
| full numerical rank? | no |
| singular value gap | 1.91875e+12 |
| singular values (MAT file): | click here |
| SVD method used: | s = svd (full (A)) |
| status: | ok |
For a description of the statistics displayed above, click here.
Maintained by Tim Davis, last updated 12-Mar-2014.
Matrix pictures by cspy, a MATLAB function in the CSparse package.
Matrix graphs by Yifan Hu, AT&T Labs Visualization Group.