Matrix: JGD_Homology/n3c6-b7
Description: Simplicial complexes from Homology from Volkmar Welker.
 |
| (undirected graph drawing) |
 |
| Matrix properties | |
| number of rows | 6,435 |
| number of columns | 6,435 |
| nonzeros | 51,480 |
| structural full rank? | yes |
| structural rank | 6,435 |
| # of blocks from dmperm | 1 |
| # strongly connected comp. | 1 |
| explicit zero entries | 0 |
| nonzero pattern symmetry | symmetric |
| numeric value symmetry | 53% |
| type | integer |
| structure | unsymmetric |
| Cholesky candidate? | no |
| positive definite? | no |
| author | V. Welker |
| editor | J.-G. Dumas |
| date | 2008 |
| kind | combinatorial problem |
| 2D/3D problem? | no |
Notes:
Simplicial complexes from Homology from Volkmar Welker.
From Jean-Guillaume Dumas' Sparse Integer Matrix Collection,
http://ljk.imag.fr/membres/Jean-Guillaume.Dumas/simc.html
http://www.mathematik.uni-marburg.de/~welker/
Filename in JGD collection: Homology/n3c6.b7.6435x6435.sms
| Ordering statistics: | result |
| nnz(chol(P*(A+A'+s*I)*P')) with AMD | 4,246,625 |
| Cholesky flop count | 7.7e+09 |
| nnz(L+U), no partial pivoting, with AMD | 8,486,815 |
| nnz(V) for QR, upper bound nnz(L) for LU, with COLAMD | 12,659,868 |
| nnz(R) for QR, upper bound nnz(U) for LU, with COLAMD | 14,851,943 |
| SVD-based statistics: | |
| norm(A) | 3.87298 |
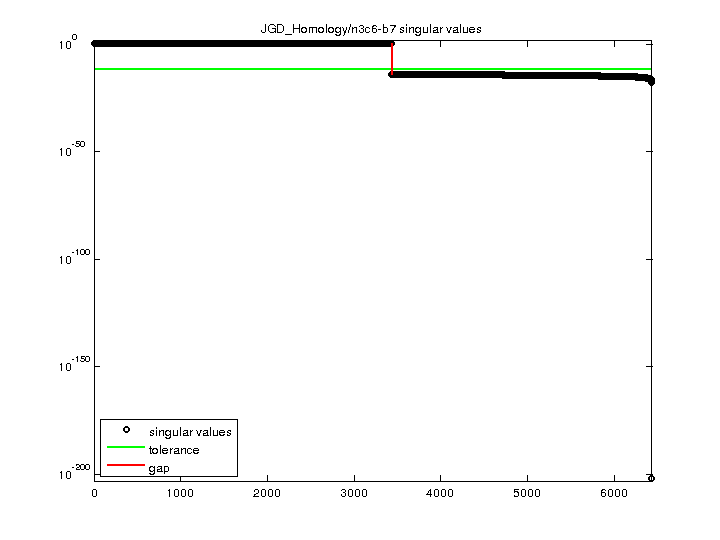
| min(svd(A)) | 7.75647e-203 |
| cond(A) | 4.99323e+202 |
| rank(A) | 3,432 |
| sprank(A)-rank(A) | 3,003 |
| null space dimension | 3,003 |
| full numerical rank? | no |
| singular value gap | 2.73097e+14 |
| singular values (MAT file): | click here |
| SVD method used: | s = svd (full (A)) ; |
| status: | ok |
For a description of the statistics displayed above, click here.
Maintained by Tim Davis, last updated 12-Mar-2014.
Matrix pictures by cspy, a MATLAB function in the CSparse package.
Matrix graphs by Yifan Hu, AT&T Labs Visualization Group.